The ancestors of cetaceans (whales, dolphins, and porpoises) moved from land to the sea over 50 million years ago. Many viruses infect cetaceans, but how they evolved during the shift from land to sea is unknown. Fossilized retroviral genomes integrated into cetacean DNA provide insight into this question.
The retroviral reproduction cycle involves the conversion of viral RNA into DNA followed by integration in host chromosomes, leading to what is know as proviral DNA. If this integration event occurs in germ cells, the provirus may be transmitted to offspring for many generations. Analysis of cetacean endogenous retroviral DNA (ERV) provides insight into when in evolution these viruses were acquired.
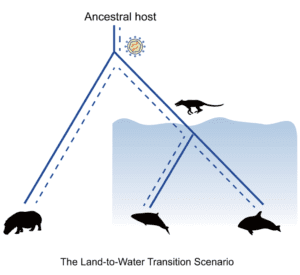
Cetacean retroviruses may have arisen in two ways. In the land-to-water scenario (pictured), the terrestrial ancestor of cetaceans was infected, and then carried the retrovirus into the oceans. In the secondary host switching scenario, only after cetaceans were in the oceans were they infected by cross-species transmission from other non-cetacean mammals.
Analysis of cetacean ERVs provides support for both scenarios. A search of the sequenced genomes of mysticetes (baleen whales) and odontocetes (toothed whales) revealed 8,724 ERVs. Phylogenetic analysis of these ERVs places them into 315 distinct lineages. Of these, 298 are found in both mysticetes and odontocetes, implying that they were present before these lineages diverged. The copy numbers of these ERVs are very low, suggesting that they did not proliferate after their hosts entered the waters. Some are related to ERVs found in hippopotamuses, which share a common ancestor with cetaceans, and are even found the the same location in the genomes.
Another 17 lineages were not common to mysticetes and odontocetes, but are found in specific sub-lineages of cetaceans. This observation suggests that they were acquired from other non-cetacean mammals. In support of this hypothesis, it was found that these ERVs are closely to ERVs in a variety of land mammals including bats and cows. These ERVs are more numerous in the cetacean genome, implying that they entered recently and might still be infecting these mammals. Whether or not the infections have pathogenic consequences is unknown.
The reader might be asking how a whale might acquire a virus from terrestrial and semi-aquatic mammals. Apparently there is more interaction of cetaceans with these animals than we think: killer whales have been observed feeding on terrestrial mammals and seals.
Given the difficulty in sampling cetaceans, it will be difficult to determine how other viruses originated in these animals. This question can be readily addressed for endogenous retroviruses because they are integrated into germline DNA, often for millions of years.

Pingback: A whale of a virus story - Virology Hub
Whales and porpoises get washed up on beaches all the time, so sampling their DNA should be relatively easy, if a bit messy. Of equal inerest is the composition of ERVs in mermaids (uh, mermen, or now maybe mermits). These are very rarely found, but they have been recently documented.
its so interesting that there might be an evolutionary relationship between whales and viruses. I support Benjamin’s comment sampling is easy for washed offshore whales
It would seem likely that cetaceans would catch viruses from effluent run off into rivers, much as the sea otters are said to have picked up infections from cats–as mentioned in early TWiPs, I think, before it became email dominated. I bet there are a lot of viruses in a cow pat, and cows are not fussy about pooing and drinking at the same time. Imagine the amount of excrement that must have got into rivers during massive buffalo, and other, herd migrations, before humans got rid of most of the mega fauna. Then there must have been vastly increased opportunity, for bovine virus spread, once humans began animal husbandry and polluted all the rivers until recently.