
I’m working on an MPH and in one of my classes we are currently studying the influenza virus. I’d forgotten that the genome is in 8 separate parts. Curious, I’ve been searching but can’t find any information as to why that is?
What evolutionary advantage is conferred by having a segmented genome?
Terrific question! Here is my reply:
It’s always hard to have answers to ‘why’ questions such as yours. We answer these questions from a human-centric view of what viruses ‘need’. We might not be right. But I’d guess there are at least two important advantages of having a segmented RNA genome.
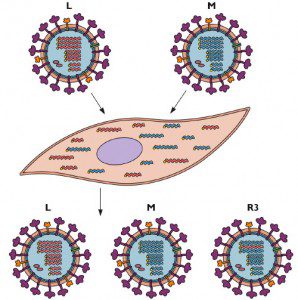
Mutation is an important source of RNA virus diversity that is made possible by the error-prone nature of RNA synthesis. Viruses with segmented genome have another mechanism for generating diversity: reassortment (illustrated).
An example of the evolutionary importance of reassortment is the exchange of RNA segments between mammalian and avian influenza viruses that give rise to pandemic influenza. The 2009 H1N1 pandemic strain is a reassortant of avian, human, and swine influenza viruses.
Having a segmented genome is another way to get around the limitation that eukaryotic mRNAs can only encode one protein. Viruses with segmented RNA genomes can produce at least one protein per segment, sometimes more. There are other ways to overcome this limitation – for example by encoding a polyprotein (picornaviruses), or producing subgenomic RNAs (paramyxoviruses).
Other segmented viral genomes include those of reoviruses, arenaviruses, and bunyaviruses.
There are various ways to achieve genetic variation and gene expression, and viruses explore all aspects of this space.

I remember having this self-same discussion back in the 1970s, when I was doing an MSc: my professor at the time waxed philosophical about how it gave an advantage in terms of length of message (=problems with copy-editing); potential for reassortment; speed of replication…. All potentially true, in retrospect. However: why are there perfectly good and successful (-) sense viruses with single-component genomes, like rabies and mumps, then? Why do totiviruses have single-component dsRNA genomes when reoviruses need 10-12 components? Why do picornaviruses have 1 ss(+)RNA while the evolutionarily-related comoviruses 2; tobamoviruses 1 and bromoviruses 3?
Because they all work, is why: each of those solutions to “how to keep a genome replicating” WORKS, is successful – so there are no advantages necessarily to ANY strategy, any more than we are more successful than a jellyfish.
Just different B-)
That’s what I tell my students – all the viral genome strategies exist because they work. But they are also evolutionarily competitive – it the niche of the particular virus, they have been successful. In my view, all viruses should have plus strand RNA genomes!
Agreed – I used to use much the same words! But I differ in that I think that circular ssDNA is especially cool.
Although lately I’m trending towards those huuuuge dsDNA guys – especially after we found a likely one in Antarctic waters B-)
My answer would be “Why not?”
I see +ssRNA, -ssRNA and dsRNA genomes as all part of the same process of RNA replication, just depends which has evolved to be put in the virion. (not quite the segmented genome issue)
The Yanagi group in Japan managed to segment the naturally nonsegmented measles virus into three segments and the recovered viruses were viable in vitro. This was possible in the lab because the molecular biology of replication is well characterised and that measles virus is polyploid allowing the engineering of a tri-segmented genome that can replicate and be packaged into virions. I speculate that this hasn’t commonly occurred in nature because measles and paramyxoviruses notoriously do not undergo recombination. However, the biological and evolutionary consequences of giving measles virus a tri-segmented have not really been explored but would shed light on this process. This question can – and should – be addressed experimentally!
Yanagi group paper: http://jvi.asm.org/content/80/9/4242.long
Pingback: Virology question of the week: why a segmented ...
I think beside the benefit of genetic reassortment, having a segmented genome in influenza would probably mean all 8 segments can be made into proteins at the same time, shortening replication time. NS1 protein is known to be expressed to high abundance early during infection, this is consistent with the fact that NS1 is encoded from the shortest segment of all.
why not recombination instead of reassortment ?
You could have the same effect by marking
some places especially determined for recombination.
And “remember” places that were successful in the past,
give them higher probability for recombination
Hi Ed,
They may all work, but the ease with which you can get a working solution is not the same. For influenza, the cost of genome segmentation is the requirement to evolve a specific packaging mechanism that allows one copy of each segment to be packaged into each infectious particle. I am not sure that we really understand how influenza accomplishes this yet (just being a casual observer, y’know). This requirement may explain why segmented measles virus genomes have not yet emerged in nature.
In plants, the high MOI of transmitted infections due to transport by insect vectors makes multipartite genomes possible, so plant viruses don’t have to go to the trouble of working out a way to package segmented genomes. This strategy does not (and will never) work in animals, or in bacteriophages, because the transmission mechanisms are different.
I think a lot of this has to do with lucky accidents: the potential set of solutions to the problem of “how to efficiently express and replicate a genome” is HUUUUUUGE, and we see SOME of the solutions that work.
Geminiviruses recombine better than rabbits do – and they are sscircDNA, like PhiX174. They have VERY distinct recombinational hotspots AND coldspots. And some have mroe than one component as well.
I think the simple and scientific answer is “Dude! Whatever works!”
I have taught for years that segmented genomes CAN allow differential expression of specific proteins, depending on the context(s) for transcription and then translation (in the case of flu), as well as for translation and then transcription, for ssRNA(+) viruses. Without requiring internal RNA promoters.
PS: good discussion group, this!
i want to ask what will happen in your body if you keep swallowing undigested food for more than 10days
in influenza-A we have the antigenically relevant HA and NA segments with multiple subtypes,
while there is virtually no evolution in the amino acids of other segments in wild waterfowl.
So the different HAs and NAs want to swap to evade immunity, but the other segments needn’t
or multiple segments are better to achieve filamentous morphology
(for whatever what that is needed)
A colleague of mine who is a well know evolutionary virologist sent me the following answer to this question:
One additional force that may drive the evolution of segmented genomes in rapidly evolving viruses (although less well commented on) might be the need to mitigate the costs of high mutation rates. For an unsegmented 7 kb viral genome at a given (high) mutation rate, the likelihood that a deleterious mutation is not encountered per 7 kb genome is modest (polio viruses for example). Thus, the greater a linear genome size, the higher the cost of linkage. Now if the same 7 kb genome was split in multiple segments, assortment allows both adaptive mutations to be propagated; because they are not always paired with deleterious mutations in a 7 kb genome, they have a higher chance of success. Thus, assortment may augment both adaptation and (significantly) mitigate the cost of high mutation rates.
I wonder if segmented paramyxoviruses would be competitive.
well, once 2 viruses infect the same cell, all the segments
float around and allow the 256 combinations.
Not so with recombination.
Pingback: Origin of segmented RNA virus genomes
Segmented rna encode different proteins required proteins with adaptable mutation.
For this free assortment is must
why do viruses have a negative sense mRNA?
Is there are convention for numbering the segments of a segmented viral genome? Are they simply numbered in order of size? The influenza A genome for example has 8 segments, which are numbered 1 to 8. In influenza A, segment 4 encodes the HA protein, while in infectious salmon anaemia virus (another orthomyxovirus with 8 genome segments) the HA protein is encoded by segment 6. Why not give the homologous segments the same number?