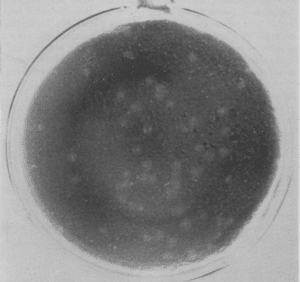

Since the early 1920s bacteriologists had used the plaque assay to quantify the number of infectious bacteriophages (viruses that infect bacteria). Dulbecco noted in 1952 that “research on the growth characteristics and genetic properties of animal viruses has stood greatly in need of improved quantitative techniques, such as those used in the related field of bacteriophage studies.” One limiting factor was the development of suitable animal cell cultures that could be used to determine viral titer. By the 1950s the techniques for reliably producing and propagating human cell cultures were developed, and in 1951 the first immortal human cell line, HeLa, was isolated. Dulbecco took advantage of these advances and showed in 1952 that western equine encephalitis virus formed plaques on monolayers of chicken embryo fibroblasts (figure). Dulbecco also made the important observation that one virus particle is sufficient to produce one plaque. He drew this conclusion from his observation of a linear dependence of the number of plaques on virus concentration. This seminal advance made possible the application of genetic techniques to the study of animal viruses.
Dulbecco’s work on tumor viruses was focused on polyomaviruses – small DNA-containing viruses such as murine polyomavirus and SV40. He found that cells from the natural host of the virus – mice for polyomavirus and monkeys for SV40 – were killed as the viruses replicated and produced new viral progeny. However, these viruses did not replicate in or kill cells from other animals. For example, when hamster cells were infected with murine polyomavirus, no viral replication took place, the cells survived, and a few rare cell were transformed – their growth properties in culture were altered and they induced tumors when injected into hamsters. Dulbecco later found that the polyomaviral DNA is a circular, double-stranded molecule; and that in non-permissive cells (in which the virus does not replicate) the viral DNA became integrated into the host cell chromosome. He also suspected that a viral protein called T (for tumor) antigen was a key to cell transformation.
Today we understand why polyomaviruses transform cells in which they do not replicate: infection does not kill these cells, and the rare transformed cells contain only viral DNA encoding T antigen. This protein is needed for viral replication in permissive cells because it drives cell proliferation, activating cellular DNA replication systems that are required for producing more viral DNA. In a non-permissive cell, T antigen drives the cell to divide endlessly, immortalizing it and allowing the accumulation of mutations in the cell genome that make the cells tumorigenic.
While the details of how DNA tumor viruses transform cells were being elucidated, other investigators were attempting to understand how another class of viruses – with RNA genomes – had similar effects on cells. In 1951 a young scientist named Howard Temin joined Dulbecco’s laboratory to study how Rous sarcoma virus (RSV) caused tumors. This virus had been discovered by Peyton Rous in 1911, but would only cause tumors in chickens, limiting progress. In Dulbecco’s laboratory, Temin found that RSV induced transformation of cultured chicken embryo fibroblasts – the same types of cells that were being used to develop the plaque assay for animal viruses. Temin took this transformation assay to his own laboratory, where he reasoned that a DNA copy of the RSV viral genome must be integrated into the chromosome of transformed cells. This led him to discover the enzyme reverse transcriptase in RSV particles, which produces a DNA copy of the viral RNA.
By embracing a new technology for the study of animal viruses – cell culture – Dulbecco set the study of both DNA and RNA tumor viruses on a path that would lead to understanding viral transformation, an achievement recognized by the 1975 Nobel Prize.
Dulbecco, R. (1952). Production of Plaques in Monolayer Tissue Cultures by Single Particles of an Animal Virus Proceedings of the National Academy of Sciences, 38 (8), 747-752 DOI: 10.1073/pnas.38.8.747

And only a few days ago you guys were talking about one of his papers!
I find it amazing that Dulbecco was publishing work up to only a few years ago on breast cancer – truly inspirational and not only to virologists.
…and he co-authored a very influential textbook on Virology – which was my bible for quite a few years.
We need to get some more history from these pioneers before they all disappear!
Did he also create Dulbecco’s Modified Eagle’s Medium (DMEM)?
Apparently so, http://www.atlantabio.com/catalog/media-&-balanced-salts/classical-media/dmem
Indeed, he did develop DMEM, and we use it all the time.
I hope I last as long. If I did, TWiV would be up to #2072!
Thank you for this very nice profile.
Pingback: TWiV 179: Was ist ein virus?
Pingback: TWiV 179: Was ist ein virus? | Alan Dove, Ph.D.
Pingback: 250 Angstroms. Keep your books. | California Southern