
Readers of virology blog know that in 2009 Lombardi et al. published a Science report indicating they had detected the new retrovirus XMRV €“ first detected a few years earlier in prostate tumors €“ in the blood of a high proportion of patients with chronic fatigue syndrome. Many other laboratories attempted to reproduce this finding, but none were successful.
The next year Alter and colleagues reported finding retroviral sequences in the blood of a substantial number of CFS patients. No viruses were isolated in the Alter study; viral sequences were obtained by polymerase chain reaction (PCR). The viral sequences were not XMRV, but were closely related to endogenous retroviruses of mice called polytropic murine leukemia viruses. (Polytropic means the viruses can infect many species, including mice; xenotropic means that the viruses, though originating in mice, only infect non-mouse species).
The Lo-Alter finding was viewed by many (including myself) as supporting the findings of Lombardi et al., but upon closer inspection it became apparent that they only clouded the situation. The viral sequences reported in the Alter study were not XMRV, and it was not clear why CFS would be caused by such a diverse range of viruses. A second report in 2011 reported MLV-like sequences in a CFS cohort but many other studies failed to find any kind of retrovirus in the blood of CFS patients.
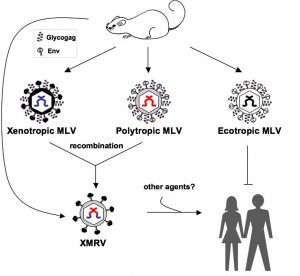
Earlier this year it became clear that XMRV is a laboratory-generated recombinant murine retrovirus: it arose during the passage of a prostate tumor in nude mice in the early 1990s. This finding made it highly unlikely that the virus could be associated with human disease. Lombardi and colleagues then retracted part of the 2009 Science paper that reported viral nucleic acid sequence; they noted that their samples were contaminated with XMRV plasmids. What remained of the paper were serological and virus culture experiments that were not specific for XMRV. Last week the remainder of this paper was editorially retracted by Science.
That left the Lo-Alter findings. The first warning came from the observation made by several laboratories that reagents used to carry out PCR are often contaminated with mouse DNA (an example is Singh’s study). The presence of this adventitious DNA can lead to detection of MLV-like sequences that resemble those found in the Lo-Alter study. The implication was clear: the Lo-Alter findings were wrong, a result of contamination of PCR reagents with mouse DNA.
More doubt came from a report of the Blood XMRV Scientific Working Group, which was assembled to determine if XMRV constituted a threat to the blood supply. In this study, sets of coded samples previously shown to be XMRV positive, as well as samples from healthy controls, were blinded and provided to 9 laboratories for analysis by PCR, virus culture, and serology. Two laboratories reported evidence of XMRV in the coded samples. Only the Whittemore-Peterson Institute identified positive specimens by PCR: two from negative controls, and one from a CFS patient. The Lo laboratory did not detect any positives by PCR, using the same nested assay that they had previously reported in their PNAS paper. The samples tested included 5 specimens that were positive in the Lo-Alter study.
The retraction of the Lo-Alter PNAS paper curiously begins with the assertion that the authors could not detect contaminating mouse DNA in their samples €“ which was most certainly present and lead to their detection of MLV-like sequences.
Although our published findings were reproducible in our laboratory and while there has been no evidence of contamination using sensitive mouse mitochondrial DNA or IAP assays or in testing coded panels…
This failure remains puzzling and unexplained; but as they report in the next paragraph, they appear to have run out of material to distribute to other laboratories for ‘independent confirmation’.
The authors provide three additional reasons why they are retracting this paper. They note that no one has been able to reproduce their findings, including the Blood XMRV Scientific Working Group. They have not been able to find (along with collaborators) anti-XMRV antibody, XMRV virions, or viral integration sites in patient samples. Finally, they mention their finding from the PNAS paper that a second set of samples taken 15 years later from the same CFS patients also were positive for MLV-like viruses. Phylogenetic analyses revealed that these sequences were clearly not descendants of the original strains. The sequence data used to make this conclusion were available for the PNAS publication, so it is not clear why this evolutionary incompatibility was not noted previously.
The authors conclude:
…in consideration of the aggregate data from our own laboratory and that of others, it is our current view that the association of murine gamma retroviruses with CFS has not withstood the test of time or of independent verification and that this association is now tenuous. Therefore, we retract the conclusions in our article.
The retraction of the Lombardi et al and Lo-Alter papers erases the published evidence suggesting involvement of a retrovirus with CFS. While it is theoretically possible that CFS has a viral origin, at the moment there are no data in support of a specific viral etiology. Some have suggested that gammaretroviruses related to XMRV might be involved in CFS. But I don’t see how a lab contaminant can point you in the direction of a bona fide etiologic agent. Contaminants cloud our vision, they do not improve it.
In light of these developments, the ongoing Lipkin study (sponsored by the National Institute of Allergy and Infectious Diseases, involving analysis of a coded panel of samples from 150 well-characterized and geographically diverse CFS patients and controls) seems less compelling. Many laboratories have failed to find any retrovirus in CFS patients, and the two papers central to this hypothesis have been retracted. Will results from one laboratory clear the matter up further? Whatever the Lipkin study finds, it will have to be validated by others – because we trust science, not scientists.
Update: The retraction has been published at PNAS.

How long until the prostate cancer papers are retracted? Patients will continue to (rightly) suspect bias and double standards until those papers are retracted too.
So whats going to happen now ? Is someone else going to come along and rediscover a retroviral connection with me/cfs and then steal all the glory ? or are we goign to be abandoned again as a patient population ?Â
To be honest I just want to be well again . I dont care who finds the cause I just want to be well but im rather hacked off by all this you only have to look at those most seriously affected like myself to know that there is some kind of virus involvement somewhere .
I get terrible bone pain , ive been on morphine for over 2 years why isnrt anyone looking at our bone marrow ? Ive said all along thats where the answer more than likely lies in me . Or lymph nodes . Ive had lymph nodes come up the size of half an egg , why not take biopsies and find out just what is lurking there instead of just ignoring them and then saying well its gone back down now but weve no idea why it swelled up so mcuh .
My immune system is now showing many signs of abnormalities and still im not taken seriously . Im hypothyroid with antibodies (hashimotos ) ive also tested positive on the ANA test more than once . Does it make any difference to how I am treated in the UK does it hell . My blood tests are always abnormal yet we are treated as though we have mental illness and then they wonder why CBT doesnt work for people liek me . Infact I was told at the ME/CFS clinic I hadnt tried hard enough with the CBT . yes you read that right totally laughable unless your on the recieving end .Â
So come on scientists enough trying to score points of one another , enough being told what to do by powers from above . Theres a bunch of really sick people here who really all they want is to be well again .Â
No more politics , and end to the psychobabble we get in the UK , show some compassion for your fellow mankind . ME/CFS doesnt care wether your black, Â white, Â rich , poor , young or old . Its ok you all burying your heads in the sand , and trying to palm it off as a mental illness for the past several decades but more and more people are getting sick . Next time it could be you or someone close to you . Is that what it takes before people actually help us ?
Â
So whats going to happen now ? Is someone else going to come along and rediscover a retroviral connection with me/cfs and then steal all the glory ? or are we goign to be abandoned again as a patient population ?Â
To be honest I just want to be well again . I dont care who finds the cause I just want to be well but im rather hacked off by all this you only have to look at those most seriously affected like myself to know that there is some kind of virus involvement somewhere .
I get terrible bone pain , ive been on morphine for over 2 years why isnrt anyone looking at our bone marrow ? Ive said all along thats where the answer more than likely lies in me . Or lymph nodes . Ive had lymph nodes come up the size of half an egg , why not take biopsies and find out just what is lurking there instead of just ignoring them and then saying well its gone back down now but weve no idea why it swelled up so mcuh .
My immune system is now showing many signs of abnormalities and still im not taken seriously . Im hypothyroid with antibodies (hashimotos ) ive also tested positive on the ANA test more than once . Does it make any difference to how I am treated in the UK does it hell . My blood tests are always abnormal yet we are treated as though we have mental illness and then they wonder why CBT doesnt work for people liek me . Infact I was told at the ME/CFS clinic I hadnt tried hard enough with the CBT . yes you read that right totally laughable unless your on the recieving end .Â
So come on scientists enough trying to score points of one another , enough being told what to do by powers from above . Theres a bunch of really sick people here who really all they want is to be well again .Â
No more politics , and end to the psychobabble we get in the UK , show some compassion for your fellow mankind . ME/CFS doesnt care wether your black, Â white, Â rich , poor , young or old . Its ok you all burying your heads in the sand , and trying to palm it off as a mental illness for the past several decades but more and more people are getting sick . Next time it could be you or someone close to you . Is that what it takes before people actually help us ?
Â
Tony Mach is popping up all over patient forums, telling us how to think.  His posts are often angry and often cruel.  He was banned from mecfsforums for this reason, by request of many members.  You can read his comments on treatingxmrv here http://treatingxmrv.blogspot.com/2011/12/tunnel-vision.html Â
On the same blog, a few days ago, Dr. Michael Snyderman reported on his progress on ARVs, which have given him an extra eighteen months of life so far: Â http://treatingxmrv.blogspot.com/2011/12/update-from-michael-snyderman-md.htmlI can tell you that all those asking that the research into gamma-retroviral involvement is allowed to continue are patients. Â I know Polly and Hone-5kl well. Â We are all sick with Myalgic Encephalomyelitis. Â Believe it or not if you wish. Â I am fairly sure that Tony Mach, RRM and Lance take delight in our suffering. Â To hear a young woman with ME called a troll, by a troll, is ironic.The blood working group study did not exclude patient contacts from the controls. Â There are many other questions unanswered on the methodology of that study. Â Why can our legitimate questions not be answered? Â This is our lives on hold here, unable to do much more than lie on the sofa with a laptop propped upon our knees.
The fact remains that Lo and Alter stand by their research, as do Ruscetti and Mikovits.  Is anyone here aware of the work by Grossman, who found the same virus in 1986 but failed to get it published until 1997?  He uploaded sequences of his virus to genbank in 2010: you can see it here http://www.ncbi.nlm.nih.gov/nuccore/HM119591.1
The terminology for the putative retrovirus associated with ME has not settled yet. Â XMRV/MuLV/HGRV etc are not factual names, they are temporary labels that get switched around as understanding changes. Â Enough with the semantic arguments already.
Lipkin, Mikovits and Ruscetti will show that Science magazine made a knee jerk reaction. In the end the science will prevail!
I totally agree with you, but no to forget the retraction of any other papers linking XMRV to disease (like Fischer et. al. who claimed that they had found XMRV in the respiritory tract of immunosuppressed patients). XMRV ist not a pathogen. It’s settled. But unforunatley up till now only the CFS stuff got retracted. The amount of unfounded comments at this blogpost indicates how much conspiracy theory surrounds CFS. This must stop and the only way to make this stop is to prevent anything that may only look biased.
I think it makes sense that the “retraction cascade” started with Lombardi et. al. because this papers hat serious additional flaws (including the alleged manipulation of figures, wich would be clearly more a crime than a flaw). But now the other papers (who did proper science but ran into contamination issues) have to be withdrawn also.
UHM…do you have anything to say about the content of this post or are you just here to rant on about a personal issue…some would consider that trolling..
Statement from Lipkin regarding XMRV/MLV CFS/MEÂ study: http://cii.columbia.edu/blog.htm?cid=CalAzy
‘Although frequently described as the “Lipkin Study,†it is in fact the
Alter, Bateman, Klimas, Komaroff, Levine, Lo, Mikovits, Montoya,
Peterson, Ruscetti, and Switzer study, designed by these 11
investigators to bring their best methods for case ascertainment and
characterization and state-of-the-art molecular and serological
diagnostic tools to address the question of whether a retrovirus is
associated with disease.’
‘There is criticism in some quarters that this study is unnecessary given
results obtained by other investigators with other samples. However,
the participating clinical and laboratory investigators and our team at
Columbia do not agree.
We are fully committed to completing the work
rapidly and rigorously. For those who continue to express concerns that
this study is an inappropriate use of resources in a challenging fiscal
environment, please be assured that more than 85% of the funding
associated with this initiative is invested in patient recruitment and
characterization and sample collection, archiving, and distribution.
Thus, irrespective of study outcome there will be unprecedented
opportunity to explore hypotheses other than that disease is due to XMRV
or MLV infection.’
Um, I guess you have not read the rest of the comments.  My post above responds to those.  Before I made that post, I made another, I copy it here so you can see that I have  already responded to the articleÂ
“Why would any scientist expect that they could replicate the results of a study, when they did not replicate the quantitative values of the independent variables that produced the original results? Â No research group has even attempted to replicate the independent variables used by Lombardi et al or Lo et al. Â We now have the sorry situation, Â of papers being retracted because other groups have not attempted to replicate the methodology used to produce the original results.”
Lots of hope for 2012.
Um, so maybe stupid question regarding the serology….
No one has found anything with XMRV specific antibodies, as to be expected if XMRV was created in a lab by accident (22RV1).
Those who got serology results did so with some SFFV test that is supposed to be less specific covering anything MLV related (please correct me if I am wrong), so what exactly is that reacting to & why mostly only in the patients?Â
If it isn’t a sign of some other gamma retrovirus, what is it a sign of? If it is reacting not to a virus but to some random protein patients make but controls don’t, why isn’t anyone onto hunting that down as a biomarker?
Also side note – why all the pressure to retract any ME/CFS related studies ASAP but the prostate cancer studies still stand? Shouldn’t they all be retracted if XMRV isn’t really a human pathogen? Glaring double standard much?
Why the reaction appeared specific to patients? Initially probably random chance with small sample sets. In the longer term and when samples were blinded they couldn’t tell the difference between patients and healthy controls (see Blood Working Group results) so there is no biomarker to track down.
The principle reason that the CFS papers are under more pressure to retract than the prostate cancer ones is most likely related to the amount of attention they attracted. The XMRV/CFS hypothesis put forward a new disease paradigm and simultaneously raised the spectre of an incipient epidemic poised to strike down millions through a contaminated blood supply. Claims like those get investigated and in this case shown to be false, which leads to a rapid retraction. You’re right that the prostate cancer/XMRV papers are just as likely to be wrong and should be retracted (and hopefully will be), the difference is just one of visibility.
Ironically, given some of the claims flying around, the XMRV/PMLV hypothesis didn’t die because science doesn’t care about CFS, it died because science does care and many labs attempted to repeat the findings and ultimately couldn’t. Normally it would take decades to get to this point and the original paper would be forgotten rather than retracted.
Nice response Ashley.Â
The antibody test in Lombardi et al. can only identify exogenous MLV viruses. Â Thus is can identify both poltropic and xenotropic MRV infections if present.
There is no evidence in the history of scientific research into this antibody that is cross reacts with anything else. Â
The Lo and Lombardi studies were both blinded. Â There is no contamination explanation or errors that have been found and there is no reason for retraction.
The blood working group study was not a replication study. Â Lo’s team used the failed assay from Lo et al. and not the assay proven to work. Lombardi also used new assays, not those from Lombardi et al. Â The controls in the study were also not deemed to be negative before blinding as no PBMCs were screened prior to coding. Â Therefore you can only look at the study as having no controls and the positives were positive. Â
The reason for retraction is purely political. Â Saying they had more attention is again a political reason. Â Studies are never ever retracted even if proven wrong for they contain valuable information. Â In this case neither paper has been found to be wrong. Â Saying this has been investigated when negative studies have only looked for viruses highly similar to one clone, VP62, that is not the viruses found in ME, is misleading. Â This is also a synthetic representation of what the viruses may be like and has never been found in nature.
The reference strain for the prostate caner viruses is VP42. So again all studies using VP62 have failed to look for those viruses. Â
Lance you are assuming that Hone-5kl is a patient or a scientist or both.
I would argue that you are harming the future of patients who do not have the energy or training to read  through all the amazing waste of bytes that you writewhich contains not a shred of reliable information, and is clear as day written by someone with neither scientific nor medical training.
If you have scientific evidence to support your comments now would be a good time to utilise them.Â
Except when you post under the name Mark G F. Â Yes?
Sorry RRM but you need to read that paper again. The assay used in the BWG was the failed assay from the Lo/Alter paper.
In Lo et al. the authors first attempted with a nested RT-PCR using Silvermans outer primers and the Lombardi inner primers to detect viruses.  This first experiment used high stringency conditions, which could only detect VP62 (a synthetic virus that does not exist in nature) That is the assay repeated in the BWG.
The assay that did find the viruses was the second assay.  This used the lombardi outers and a home made set of inners to select for sequence variability.  In this successful assay they dropped the annealing temperatures to 52C to increase sensitivity.
There has been no replication study of the methods used in Lombardi or Lo. Â The negative papers all looked for XMRV/VP62, not the PMRVs found. Â
Why don’t YOU conduct your own replication study and submit is for publication and peer review?Â
Their reference test is the clinically validated assay from Urisman et al. and the other experiments that prove it can only be a MRV. Â The serology and the EM are two such assays.
I am waiting for a scientific explanation Lance not your opinion. Â
Nothing has been settled. Â There is no evidence of contamination and the negative studies have been using novel unproven assays. Â Come on show one study that found contamination in the samples of Mikvoits, Rusectti, Lo or Alter? Â There is no evidence, you have built up your beliefs from what? Â A dislike for science? Â
There is no evidence of any flaws in Mikovits or Ruscetti experiments. Â There was no manipulation of figures either. Â The gels labels were altered by Science for publication to protect the patients identities and simplify the experiment for publication. Â They knew about AZA as they saw it for the peer review. Â Patients themselves have discussed the use of that for the last two years. Â
Lance the above is actually a quote from Professor Racaniello from an interview he did after the publication of the CDC paper.
http://forums.phoenixrising.me/content.php?187-Dr-Mikovits-and-Dr-Racaniello-on-XMRV
Lance you have to take the evidence  for what it is.  Originally the viruses isolated by Mikvoits and Ruscetti were said by Silverman to be XMRV.  Others then claimed they could only be VP62/XMRV.  Silverman’s data was wrong and the viruses are only polytropic. Â
The rest of your post is about CFS, which is a label, not ME.
Actually Sidney Grossberg 1997 paper on MRVs is still published. Â Pretty sure DeFraites is too.
It is very easy to test the hypothesis outlined in Lombardi and Lo, so why the refusal by scientists to do so? Â Who is telling them it is not safe to research this?
I think it is clear you have no scientific training. Â
Can we have some science please.  People are not testing for the viruses found.”A PMA may be required if the test is for a novel agent, if it is a new method for which clinical relevance and clinical use must be established, or if the analyte poses a health threat (such as Mycobacterium tuberculosis) and a false-positive or false-negative result would be of significant risk to the patient or general public. The manufacturer must provide data that demonstrate a reasonable assurance of safety and effectiveness of the device. FDA review of a PMA application includes an in-depth scientific evaluation of the data and a comprehensive good manufacturing practice inspection at the manufacturing facility.”http://cmr.asm.org/content/23/3/550.full”Clinical test validation: This section describes the ability of a test to detect or predict the associated disorder in patients and includes clinical sensitivity and specificity, and/or other performance attributes of testing biological samples.””When a new diagnostic is being considered for use in selecting patients to receive or to avoid a particular drug therapy (i.e., drug/test co-developed product) or to stratify patients in some other way, two distinct, but related, issues should be addressed.The first is the ability of the test to select (or deselect) patients with the biomarkers (analyte(s)) of interest. This is clinical test validation — use of a test to detect or predict the associated disorder of interest in biological specimens from the target patient groups. This should be considered the domain of clinical test validation.”http://www.fda.gov/downloads/drugs/scienceresearch/researchareas/pharmacogenetics/ucm116689.pdf
You’ll be waiting along time if you’re not satisfied with what is already out there I’m afraid.
I am assuming Hone-5kl is you.
The medical establishment will have you believe that Chronic Fatigue Syndrome/Myalgic Encephalomyelitis (CFS/ME) is some sort of ‘mysterious illness,’ but it’s no mystery to me; CFS/ME leads to HIV-Negative AIDS, idiopathic CD lympocytopena (ICL), a clinical diagnosis that I possess.
How can the AIDS establishment continue on with a stale “it’s caused by HIV” mantra when there are ICL cases cited in medical journals dating back to 1992? While millions of ailing immunodeficient CFS/ME patients get belittled and neglected, perfectly healthy HIV+ people are allocated billions of dollars in taxpayer money.
How can it make any sense to anyone?
It’s so easy to see that the medical establishment simply has these paradigms (CFS, HIV) inverted. {more} http://www.cfsstraighttalk.blogspot.com
Oh dear gods!
I guess you have not read the BWG paper by Simmons et al. then.
In that study, Ruscetti obtained positive antibody signals from 11 out of 15 controls (using his own validated test and without letting Vinnie fly in to do the testing). No matter how good or bad the controls were pedigreed, this is only explainable by false positive results (which means your assays are cross reacting with *something*)
In fact, even when you’d selected 15 close contact controls and you’d done NO pedigreeing AT ALL, you would not expect more  positves signals from the controls in this study than from patients in the original study.
No, Diago.
In one post at this blog, someone with rather low morals took my name “RRM” and decided to post a troll message using it.
The site’s software then decided to award the troll the screen name “RRM” and automatically changed the screen name on of my posts in that topic (the old ones as well as the posts after that) to Mark G F.Â
The reason for this choice must simply be the fact that I used (and am using) mark.g.f(at)somedomain.com as my email address when posting as RRM.
But nice investigating, Sherlock.
No.
The site’s software changed my name on that occasion to Mark G F, because a person with an apparent lack of morals decided to post a troll message using the same RRM handle.Â
Apparently, the site’s software does (or did) not accept identical user names (using different email accounts), and so it changed my screen name to Mark G F on all post that were already made and yet to be made in that comment section.
The software apparently chose this name because the email account I use when I post as RRM is mark.g.f(at)somedomain.com.
But excellent investigation, Sherlock.
”
That is the assay repeated in the BWG”
No. The annealling temperature that is mentioned in the BWG paper, is in reference to the IAP assay.Â
As Lo et al. definitely state that they’ve found patients positive using the BWG primers and they definitely show a gel that has some positive controls in it with those same primers, in conjunction with the fact that Simmons et al. states that assays were essentially performed as perviously described, it is safe to assume that they used the correct annealling temperatures.
Moreover, Lo et al. state in their original paper that:”The nested PCR results produced by the second round of amplification using either the internal primer set GAG-I-F/GAG-I-R (with a predicted size of an ∼410-bp product) or the internal primer set NP116/NP117 (with a predicted size of an ∼380-bp product) were essentially identical.”So no, there was not one primer set that “worked” and another one that didn’t. The PCR results for both primer sets were “essentially identical” according to the authors.Â
I am not a scientist, but I’m pretty sure the sample size was large enough that it would be extremely unlikely that the higher percentages in patients was chance (patients n=101)
Lombardi was blinded. It was misreported, pretty sure by Jon Cohen, that Lombardi was not blinded. That article had a lot of inaccuracies biased against ME patients and Mikovits (I assume Lo was blinded also).
Jane, sorry for the delay in replying. I don’t know if you are still reading but here goes anyway….
No one is telling you how to think. Yes Tony Mach is a little blunt in places but I have seen nothing that I would describe as “angry and cruel,” certainly no more so than what has been said to him. I would have phrased what he said what he said on JDJ’s blog a little differently but I cannot fault him for accuracy.
I take no pleasure in you (or anyone else) being ill. I am sure that RRM and Tony Mach feel the same. However, that does not extend to “I am ill therefore you must accept everything I say.” You have posted on a publicly accessible blog a view that is widely held to be wrong. Now, while I would vigorously defend your right to express your view, you cannot complain when people disagree with you. Perhaps their vehemence simply reflects the degree to which they feel you are incorrect. To write with self referential overtones implying people want you to be ill and to continue to cling to the idea that there is any link between a murine retrovirus and CFS/ME is bound to make you marginalised. If you have CFS I’m sure you feel marginalised already but you are only compounding this problem.
I am really sorry that you are ill, and I hope you get better. If it is any consolation, I do believe you have a real illness, I just don’t know what it is and why it happened. Please don’t take disagreement (especially on science blogs) personally – it is the essence of science.
Pingback: A Tale of Two Viruses: Why AIDS Was Pinned to HIV, but Chronic Fatigue Remains a Mystery | Natural Remedy For Depression
Pingback: A Tale of Two Viruses: Why AIDS Was Pinned to HIV, but Chronic Fatigue Remains a Mystery | The Crux « Science Technology Informer
What a load of crap. If it was in any way related to a retrovirus then dozens who had tried antiretroviral drugs would’ve improved. They didn’t, with the sole exceptions of Deckoff-Jones (who later recovered without them) and that dude with cancer.
Jane, the ME/CFS forums, which are thankfully gone from the internet, did more to damage our (patient) reputations than anything else in the history of the illness.
Please thank that dictator Patricia Carter and her f-buddy Robyn Erland from all of us who are trying to recover. Thanks for making us all look like total nutjobs and setting us back at least 30 years.
Thank you for sharing your opinion. Contrary to your belief, my NON HIV AIDS case goes up through the NIH, CDC, White House, WHO, to the UN. I testified federally in Washington, DC, sat on conference call with the American Red Cross, and have been flown out of the USA, twice now facilitated by the UN, to give my blood. Have a nice day.
“NON HIV AIDS CONFIRMED IN ASIA: IS All AIDS NON HIV CAUSED?”
“Today, in breaking news, NIH confirmed existence of NON HIV AIDS… Watch CNN video:.. ”
http://newsrescue.com/hiv-aids-confirmed-asia-aids-hiv-caused/#axzz2GlZkO1O4
I’m not sure what your case has to do with ME/CFS, which is definitely not contagious or transmittable. But thanks for the interesting link, and hope you’re improving w/ARVs.