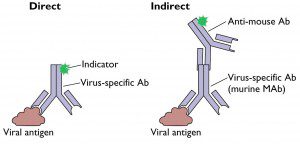

In direct immunostaining (illustrated), an antibody that recognizes a viral antigen is coupled directly to an indicator (a fluorescent dye or an enzyme). Indirect immunostaining is a more sensitive method because a second antibody is coupled to the indicator. The second antibody recognizes a common epitope on the virus-specific antibody. Multiple second antibodies can bind to the first antibody, leading to an increased signal from the indicator compared to direct immunostaining.
To carry out immunostaining, virus-infected cells are fixed to preserve cell morphology or tissue architecture. This step is usually accomplished with acetone, methanol, or paraformadehyde. After incubation of fixed cells with the appropriate antibody, excess antibody is removed by washing, followed by microscopy. Common indicators that are coupled to antibody molecules include fluorescein and rhodamine, which fluoresce on exposure of the cells to ultraviolet light. Filters are placed between the specimen and the eyepiece to remove blue and ultraviolet light; this ensures that the field is dark, except for cells that have bound antibody. These emit green (fluorescein) or red (rhodamine) light.
Antibodies can be coupled to indicators other than fluorescent molecules. Examples are enzymes such as alkaline phosphatase, horseradish peroxidase, and β-galactosidase. These enzymes can convert an added substrate to a colored dye. For example, the bacterial enzyme β-galactosidase converts the chromogenic substrate X-gal to a blue product, which can be visualized by microscopy.
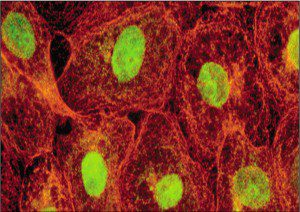

Other uses of immunostaining include monitoring the synthesis of viral proteins, determining the effects of mutation on protein production, and investigating the sites of virus replication in animal hosts. Immunostaining of viral antigens in clinical specimens is also used to diagnose viral infections. Direct and indirect immunofluorescence assays with nasal swabs or washes are routine for diagnosis of infections with respiratory syncytial virus, influenza virus, parainfluenza virus, measles virus, and adenovirus.
When cultured cells are examined by this technique it is called immunocytochemistry; when tissues are studied, the procedure is immunohistochemistry. Flow cytometry is yet another way to use immunostaining to study the synthesis of one or more proteins in cells.

Pingback: Tweets that mention Detecting viral proteins in infected cells or tissues by immunostaining -- Topsy.com
Dear Prof. Racaniello,
is there any way to predict which fixation reagent works best for which cell line? We routinely use Vero cells for the ICC but our problem is that the cells peel off during the last washing steps (before adding the substrate and after that). I remembered from growing 293 cells that coating the plates with poly-D-lysine might help the cells to stay attached but that didn’t work at all. We tried 80% methanol (-20 °C), 60%/40% methanol/acetone (-20 °C), and5%  glutaric dialdehyde solution (room temp). I also tried to modify the washing steps to reduce the physical stress on the cell monolayer (25 mL stripette, multichannel pipettor, squirt bottle) but I don’t seem to find the sweet spot for no damage. On a side note, not all wells seem to peel off though.
Any suggestions are appreciated!
Many thanks,
Sven