
There are now 64 laboratory confirmed cases of infection with the H1N1 swine influenza strain, up from 40 the day before. States reporting cases are California (10), Kansas, (2), New York City (45), Ohio (1) and Texas (6). These are the same states that reported isolations on the previous day. There are new laboratory confirmed isolations of the virus in Australia (3), Israel (1, a traveler returning from Mexico), and New Zealand (3). The number of laboratory confirmed cases in Mexico remains the same as the previous day: 26 cases, 7 deaths. This brings the total number of countries reporting laboratory confirmed cases to seven.
Swine influenza virus isolates from the US and Mexico have been given names according to the proper nomenclature, which takes the following form:
Influenza type/Country/isolate number/year (subtype).
Accordingly, the following swine influenza virus strains have been isolated:
A/California/04/2009 (H1N1)
A/California/06/2009 (H1N1)
A/California/07/2009 (H1N1)
A/California/08/2009 (H1N1)
A/California/10/2009 (H1N1)
A/Texas/04/2009 (H1N1)
A/Texas/05/2009 (H1N1)
A/Kansas/03/2009 (H1N1)
A/Ohio/07/2009 (H1N1)
A/New York/19/2009 (H1N1)
A/New York/20/2009 (H1N1)
A/Mexico/4482/2009 (H1N1)
A/Mexico/4486/2009 (H1N1)
A/Mexico/4108/2009 (H1N1)
A/Mexico/4115/2009 (H1N1)
A/Mexico/4603/2009 (H1N1)
A/Mexico/4604/2009 (H1N1)
I expect to see many more isolates from different countries in the coming weeks.
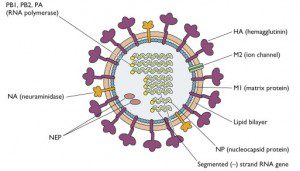
Today the CDC released genome sequences of the viral RNAs of six swine flu isolates from California and Texas (Addendum: sequences of New York, Ohio, and Kansas isolates were added late yesterday). The influenza virus genome consists of eight segments of RNA, each coding for one or more proteins (illustrated). Each RNA segment has a name – PB2, PB1, PA, HA, NP, NA, MP, and NS. Mystery Rays has done a quick analysis of the sequences. The isolates are all the same strain, but they are not identical. Unfortunately we don’t yet have genome sequence from any Mexican isolate – otherwise we could determine if they are significantly different. Such information might provide clues about why the disease in Mexico seems to be more severe than elsewhere.
A comparison with RNA sequences of other influenza virus isolates shows that most of the viral RNAs are from swine influenza viruses, with the possible exception of the PB1 RNA, which may be derived from a human H1N1 virus. This observation is somewhat surprising, because last week we were told that the new swine virus had RNA segments from pig, human, and avian influenza viruses. According to ProMED-mail, the NA and MP genes are related to those of influenza viruses from Asian-European swine, and the other genes appear to originate from swine flu viruses from pigs in North America. The data are in accord with the original assertion of the CDC that all genes of the new isolate were derived from swine viruses.
The fact that A/California/04/2009 and related isolates are pig viruses, with little or no genetic material from human influenza virus strains, is fascinating. Clearly these strains are different from viruses that circulate in pigs because they can be transmitted among humans and cause respiratory disease. It will be very important to compare the sequences of these isolates with viruses obtained from pigs in an attempt to determine what changes enabled the virus to adapt to humans.

Is it true there are zero avian flu components in these US isolates?
Yes, according to the sequences that have been submitted so far.
Pingback: About virustaxonomy, classification « Leabright’s Blog
The fact that so far the fatality rate is only one in USA (a child from Texas), it may be of a “drift” consequence in viral genome! I suggest CDC confirm it.
I would appreciate your thoughts on how likely you think it is that the higher incidence of hospitilization and death (confirmed cases) in Mexico are simply reflective of the fact that a lot more people have been infected there than in the U.S., rather than that the isolates circulating in Mexico are more virulent to humans than the isolates circulating in the U.S. If getting very sick or dying after exposure to this strain of H1N1 is primarily a function of a hyperactive cytokine response (which I assume is related to human genetics), then I am wondering if whether a person gets very sick or dies after exposure to an isolate is more related to the genetics of the person (i.e. the intensity of their cytokine response) than to the genetics of the isolate. If only a very small percentage of people have an immune response that is sufficient to make them very sick or kill them after exposure to the virus, then perhaps we simply have not had enough people in the U.S infected yet for the virus to “find” these hyper-responsive individuals.
PAN(ACA)DEMIC FLU?
*
Taking into account the 30ooo deaths per year of 'regular' flu,
how much did change this statistics the previous known epidemics (57, 68 and 86)?
*
Even if is just a 'false' alarm,
maybe such a threat boosts the knowledge about the biology and the geographical behavior of the disease,
in the same way as a vaccine boosts the human immune system,
We simply don't have enough data to know if the mortality is the same
or greater than epidemic or pandemic flu. It will take a good deal of
time to get those numbers. And yes, it's a great opportunity to better
understand the disease and its spread.
I don't see a time for this posting. The swine flu page of the CDC listed 91 cases and 1 death 11am this morning. I understand the main page still had yesterday's data.
I heard the baby was from Mexico and flown to Texas. Still doesn't seem to me anywhere near enough time or cases to estimate cfr.
I would like to know the details on structure of the virus, environmental stability, infectious dose & the disinfection. I am trying to develop risk assessment document for experimental studies on this virus. Please advice.
I assume that the above said criteria are similar to Influenza A viruses. Please advice.
The ilustration you share on this swine influenza update (showing the structure and location of the HA, NA, etc.), does it correspond to the A/California/04/2009 variety alone or is it representative of all the Influenza viruses causing the so calle “swine flue” right now?
That illustration is representative of all influenza viruses of the A
type, which includes the current swine flu, and human strains such as
H1N1, H3N2, and H2N2.
Thank you so much!
Thank you so much!
Looks like you’ve done your research very well.